Author: Chris
-
MacVector 18.2 is out! …and ready for macOS Monterey
Overview We are very pleased to announce that MacVector 18.2 is available to download. MacVector 18.2 is a Universal Binary that runs natively on both Apple Silicon and Intel Macs. It is fully supported on macOS Sierra (10.12) to macOS Monterey (12). New features: Align to Reference Enhancements Context Sensitive Hamburger Menus Where a window has…
-
Compare a pair of genomes
In recent years there has been an explosion of whole-genome sequencing projects. One common question coming out of this has been to ask: “Exactly what are the genetic differences between my sequenced organism and another related strain?” MacVector to the rescue! MacVector’s Compare Genomes By Feature… tool lets you see the differences between two annotated…
-
MacVectorTip: Scan For… Missing Primers: Automatically display Primer Binding Site on your sequences
MacVector’s Scan DNA For.. tool allows you to automatically display restriction enzyme recognition sites, putative ORFs, CRISPR PAM sites, missing annotation and also it will display primer binding sites from your own Primer Database in each DNA sequence that you open. Here’s an example of a couple of primers displayed on the pET 47b LIC…
-
MacVectorTip: Using the Align to Reference Shading and Trimming toolbar buttons
MacVector’s Align to Reference Editor and the Contig Editor in Assembly Projects have two useful functions for visualizing assemblies. The Shading button turns on background coloring of the residues in the upper pane, based on quality values (these can be from Sanger reads or from NGS reads). The scale ranges from a dark red for…
-
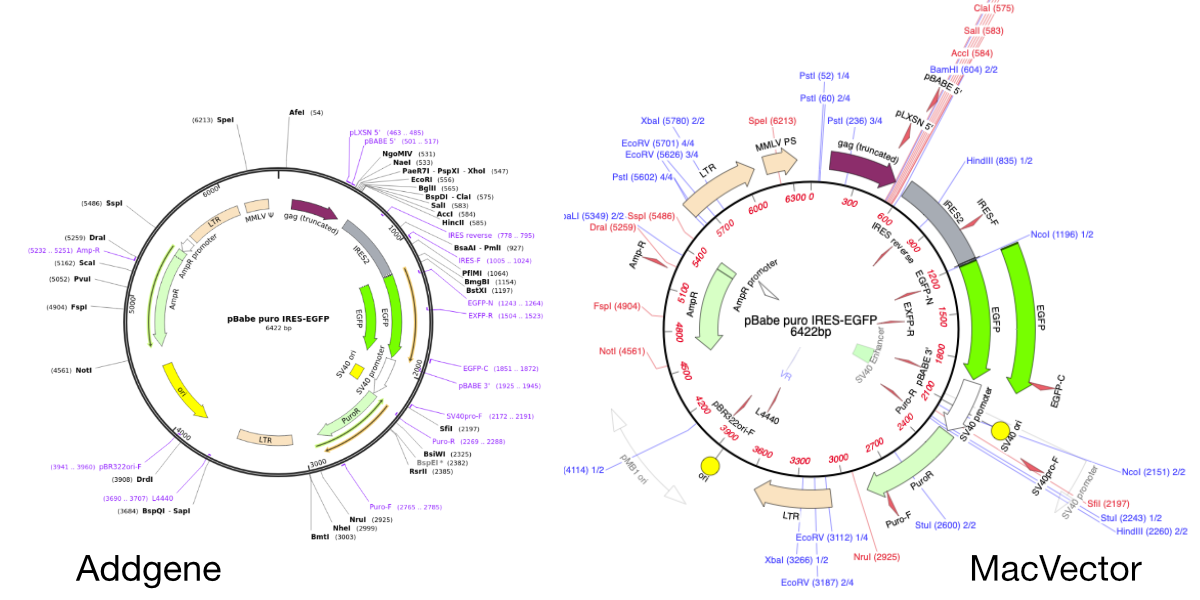
Importing SnapGene files into MacVector
MacVector will directly import SnapGene DNA files. You just need to use FILE | OPEN or double click the file. This is very useful when downloading plasmid sequences from the wonderful Addgene plasmid repository. Here’s a plasmid sequence downloaded from the Addgene website in Snapgene format. It’s been opened directly in MacVector by double clicking…
-
Designing primers and documenting In-Fusion Cloning with MacVector
The In-Fusion Cloning kits from Takara allow you to perform ligase free cloning of PCR products into vectors in as little as 15 minutes. You can use MacVector’s Gibson Cloning/Ligase Independent tool to design primers for In-Fusion cloning workflows. The In-Fusion kits need a 15nt overlap between the ends of a fragment and the ends…
-
MacVectorTip: Using tabbed sequence windows in MacVector
One of the lesser known features of macOS is the ability to store all open documents of an application in tabs. Tabs were initially introduced for the Finder, but macOS Mavericks saw them apply to supported application document windows too. MacVector has supported tabs since their introduction, however, by default the Tab Bar is turned…
-
MacVector Tip: a complex subsequence pattern example.
MacVector’s Subsequence tools allows you to search for motifs in both protein and DNA sequences. As well as a library of existing subsequence files, such as promotors and transcription factor binding sites, you can keep a library of your own subsequence matches. Subsequences libraries are multiple patterns kept in a single file. A search will…
-
Working from home and Roaming Network licenses
The pandemic brought a sudden change to usual working routines and it is probable that home working will remain part of the working week for some time to come. Most scientific research needs physical lab time, but that’s just “pipetting”! The real science also happens when you think.. and that can be done easily at…
-
MacVectorTip: Identifying, Selecting and Assembling NGS reads with a variant genotype
When analyzing/assembling/aligning NGS data, there are many scenarios where you might want to separate out the reads representing different genotypes or variant sequences. MacVector makes this very easy. Take a reference sequence and choose Analyze->Align to Reference. Now click the Add Seqs button and select and add your NGS data files. NOTE: if your reference…
